#include <Prediction.h>
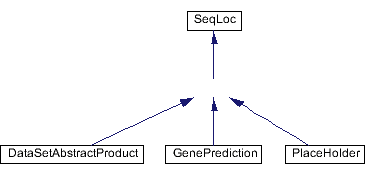
Inheritance diagram for Prediction:

Public Types | |
| typedef std::list< Prediction * > | DataStorageType |
Public Member Functions | |
| Prediction (const SeqLoc &sl, AnnotationScore d, int modelId, const string &str) | |
| Prediction (const Prediction &pred) | |
| copy constructor | |
| virtual | ~Prediction () |
| no dynamic memory management, does contain one 'stl string' object. | |
| virtual void | del ()=0 |
| AnnotationScore | getScore () const |
| return score associated with this object | |
| void | setScore (const AnnotationScore &s) |
| set score valuie | |
| int | getModelId () const |
| get integer idx of this coding sequence prediction | |
| const string & | getDataSource () const |
| get string identifier of the source of this prediction | |
| virtual DataStorageType * | makePlaceHolders () const=0 |
| all prediction type objects must be able convert themselfs into a list of pointers that contain each coding sequence seperately | |
| virtual void | print (std::ostream &os, const string &str) const |
| print to ostream, information about this prediction | |
GeneModels are a collection of GenePrediction objects.
Sequence Match models are a collection of MatchPrediction objects, both of which
are just DataSetAbstractProduct, whose behavior are outlined by this object.
In other words, one knowledge source might be made up of a collection of predictions about related sequence locations or it might be just one sequence location Presumeably, there will be many types of predictions: CDS, start or stop codons, 5' UTRs, 3' UTRS, introns, exons for starters.
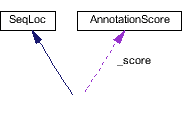
Collaborators:
SeqLoc, AnnotationScore
interface
|
||||||||||||||||||||
|
|